Mercury »
PDB 12ca-1czm »
1biq »
Mercury in PDB 1biq: Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
Enzymatic activity of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
All present enzymatic activity of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A:
1.17.4.1;
1.17.4.1;
Protein crystallography data
The structure of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A, PDB code: 1biq
was solved by
D.T.Logan,
F.Demare,
B.O.Persson,
A.Slaby,
B.M.Sjoberg,
P.Nordlund,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 25.00 / 2.05 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 74.053, 83.780, 113.953, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.1 / 26.6 |
Other elements in 1biq:
The structure of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A also contains other interesting chemical elements:
| Iron | (Fe) | 4 atoms |
Mercury Binding Sites:
Pages:
>>> Page 1 <<< Page 2, Binding sites: 11 - 16;Binding sites:
The binding sites of Mercury atom in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A (pdb code 1biq). This binding sites where shown within 5.0 Angstroms radius around Mercury atom.In total 16 binding sites of Mercury where determined in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A, PDB code: 1biq:
Jump to Mercury binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9; 10;
Mercury binding site 1 out of 16 in 1biq
Go back to
Mercury binding site 1 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
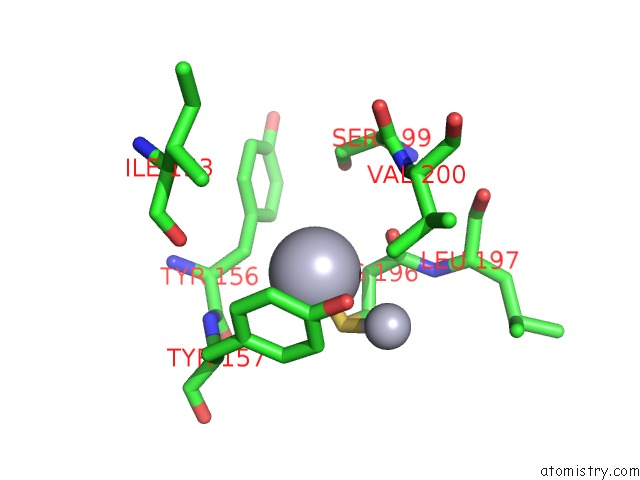
Mono view
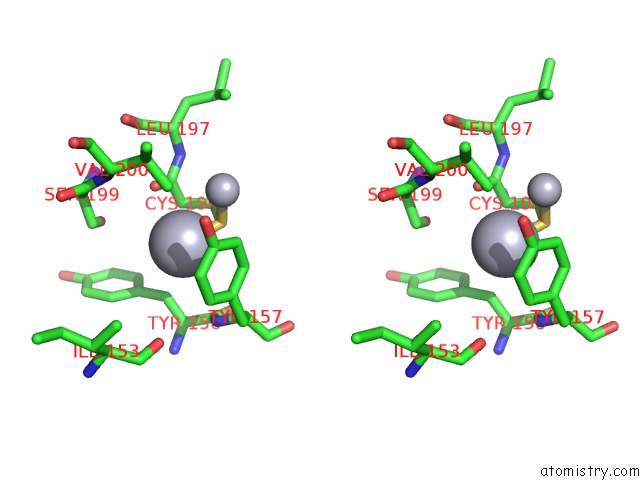
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 1 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 2 out of 16 in 1biq
Go back to
Mercury binding site 2 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
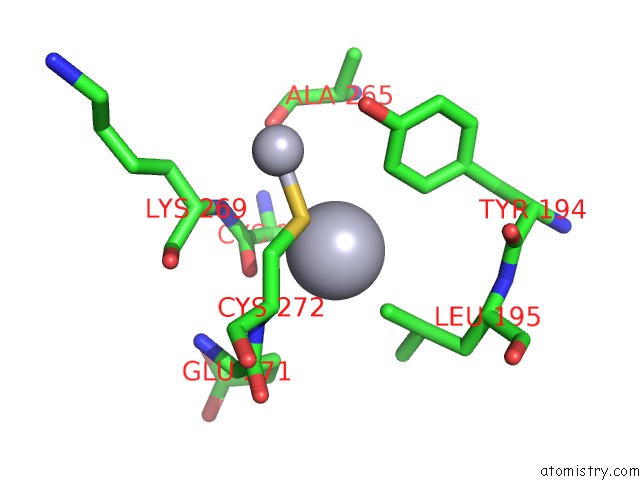
Mono view
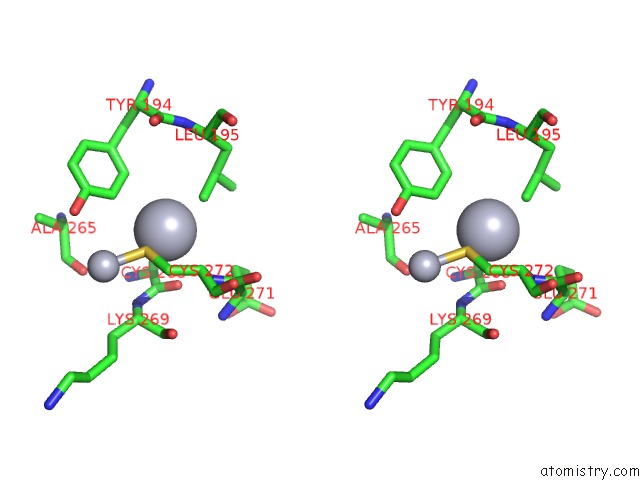
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 2 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 3 out of 16 in 1biq
Go back to
Mercury binding site 3 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
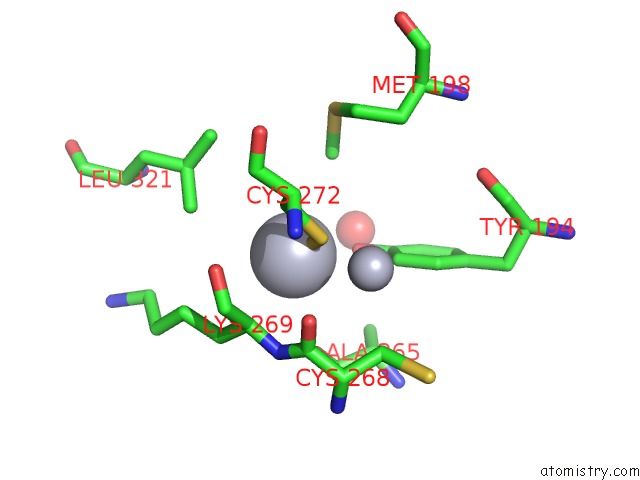
Mono view
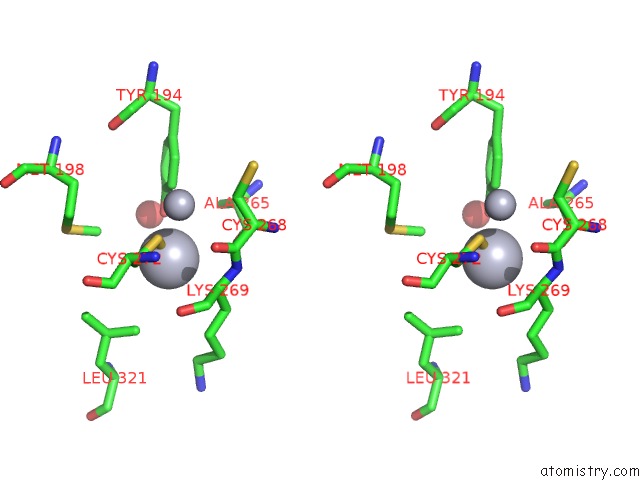
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 3 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 4 out of 16 in 1biq
Go back to
Mercury binding site 4 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
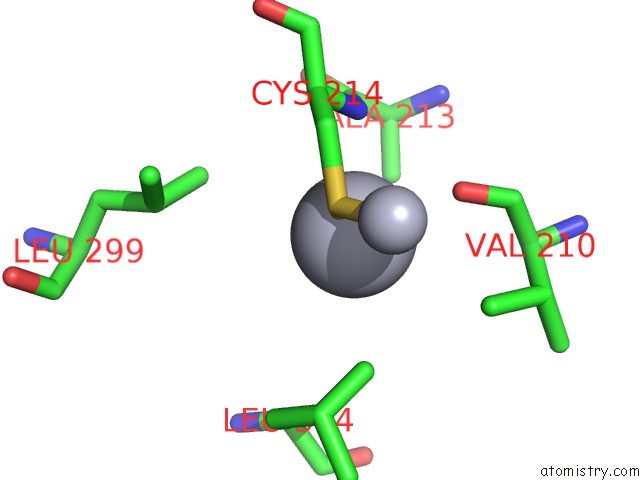
Mono view
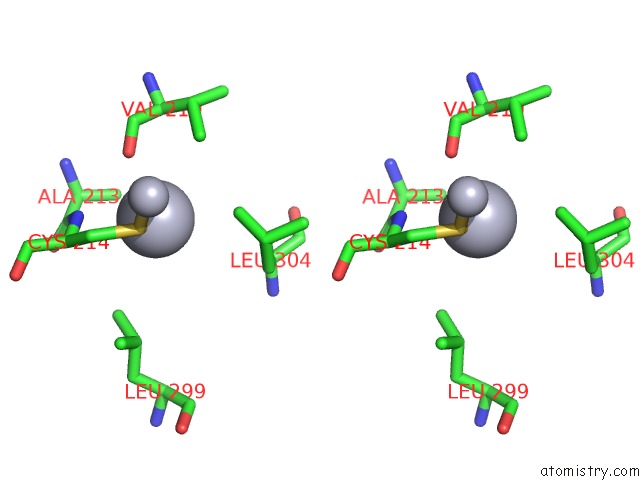
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 4 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 5 out of 16 in 1biq
Go back to
Mercury binding site 5 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
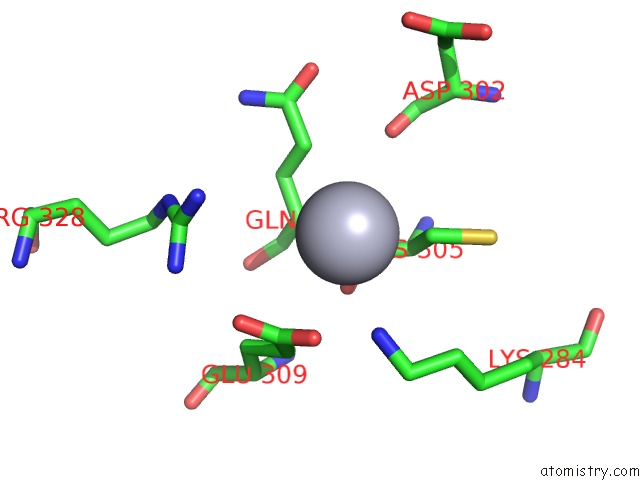
Mono view
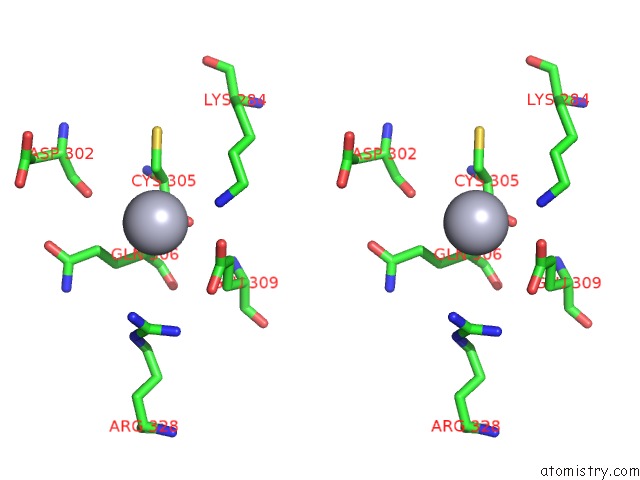
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 5 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 6 out of 16 in 1biq
Go back to
Mercury binding site 6 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
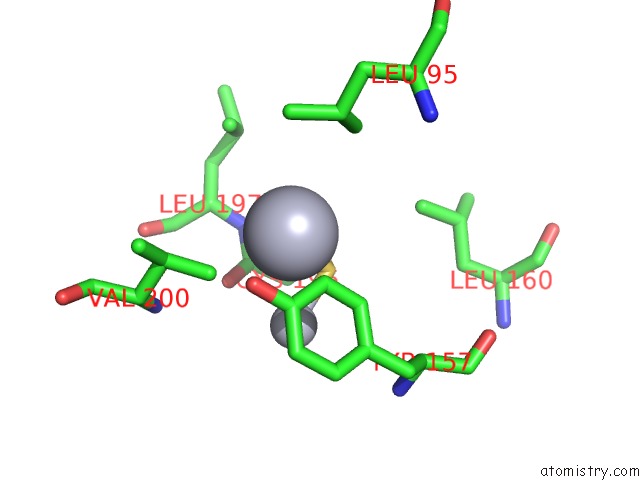
Mono view
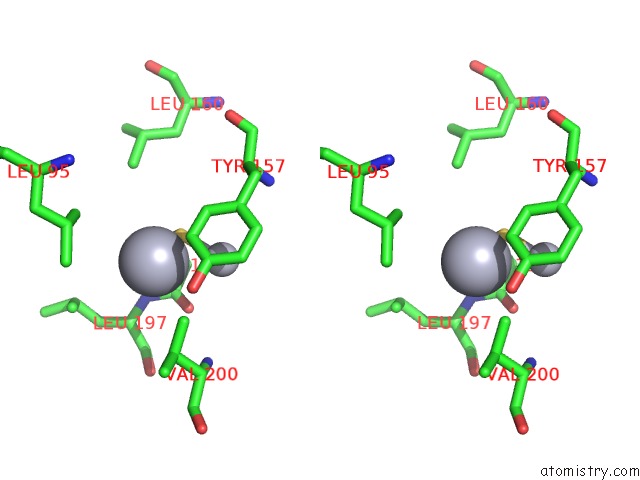
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 6 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 7 out of 16 in 1biq
Go back to
Mercury binding site 7 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
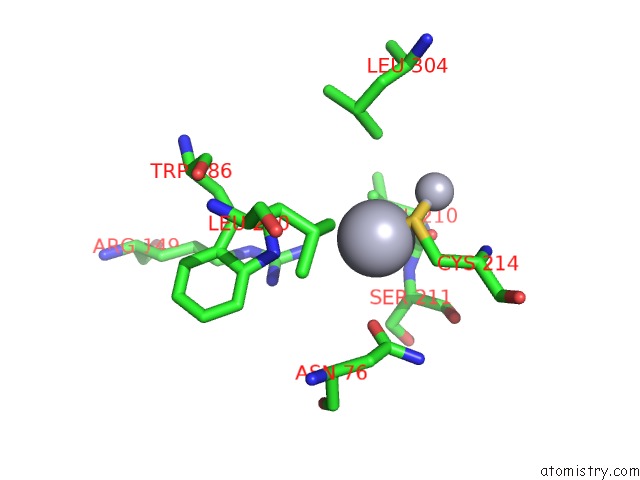
Mono view
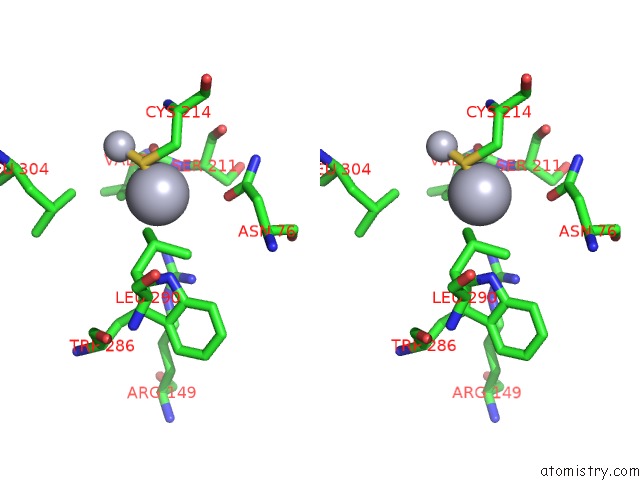
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 7 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 8 out of 16 in 1biq
Go back to
Mercury binding site 8 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
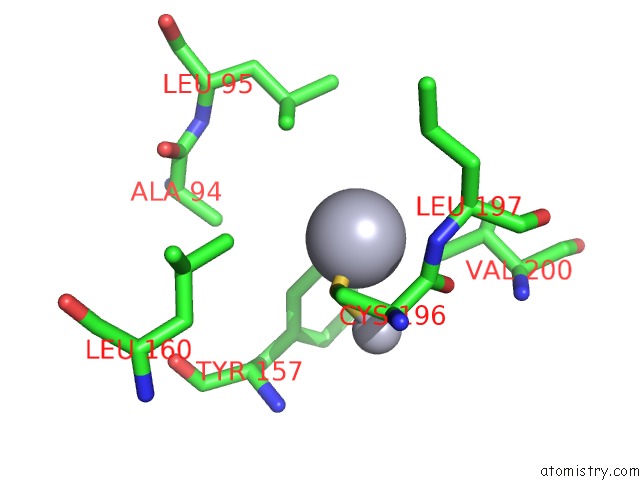
Mono view
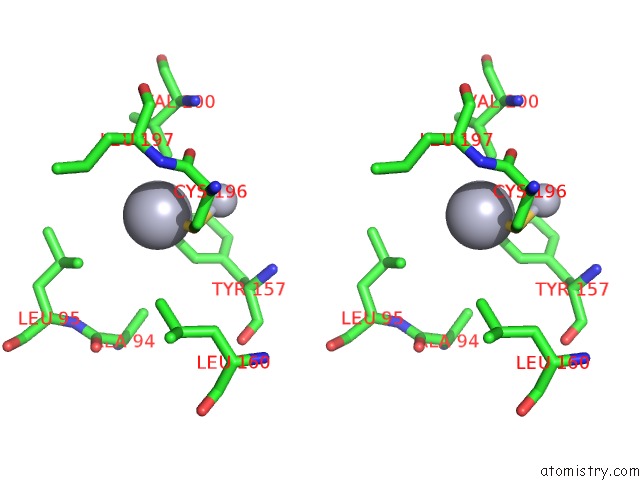
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 8 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 9 out of 16 in 1biq
Go back to
Mercury binding site 9 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
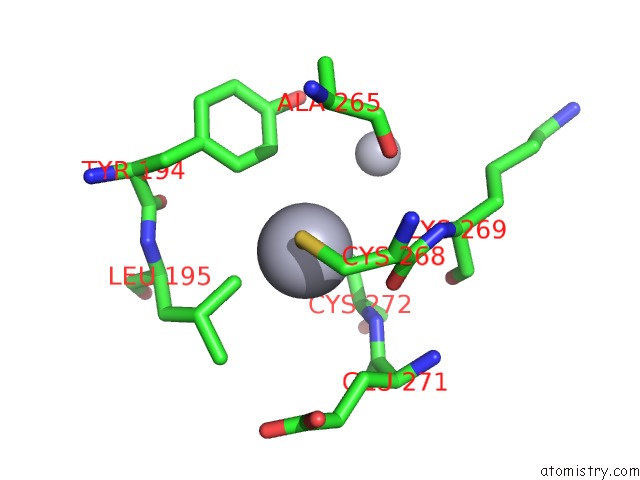
Mono view
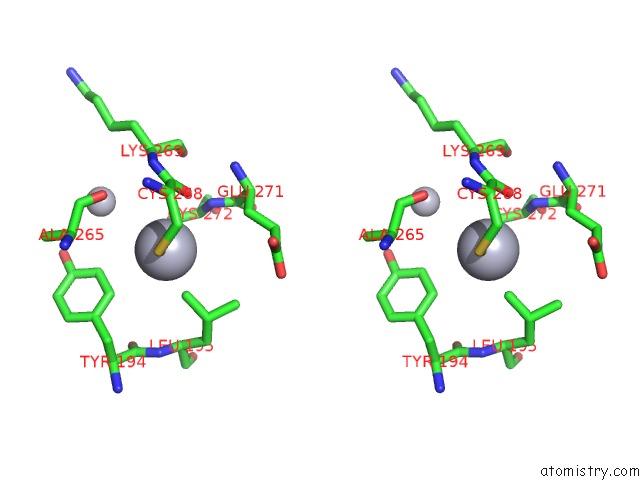
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 9 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Mercury binding site 10 out of 16 in 1biq
Go back to
Mercury binding site 10 out
of 16 in the Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A
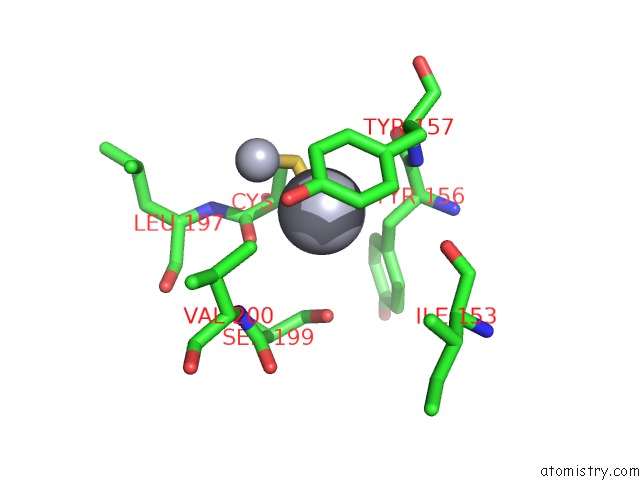
Mono view
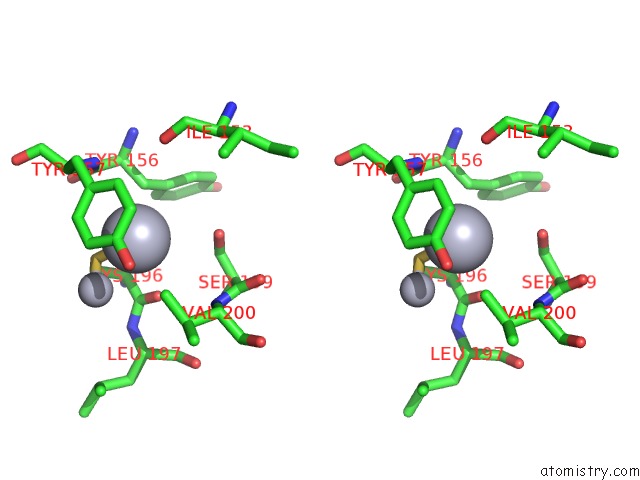
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 10 of Ribonucleoside-Diphosphate Reductase 1 Beta Chain Mutant E238A within 5.0Å range:
|
Reference:
D.T.Logan,
F.Demare,
B.O.Persson,
A.Slaby,
B.M.Sjoberg,
P.Nordlund.
Crystal Structures of Two Self-Hydroxylating Ribonucleotide Reductase Protein R2 Mutants: Structural Basis For the Oxygen-Insertion Step of Hydroxylation Reactions Catalyzed By Diiron Proteins. Biochemistry V. 37 10798 1998.
ISSN: ISSN 0006-2960
PubMed: 9692970
DOI: 10.1021/BI9806403
Page generated: Fri Aug 8 08:53:33 2025
ISSN: ISSN 0006-2960
PubMed: 9692970
DOI: 10.1021/BI9806403
Last articles
I in 3LR5I in 3LR0
I in 3KXN
I in 3LA9
I in 3LB1
I in 3L8L
I in 3L72
I in 3KZ8
I in 3KXF
I in 3KS7