Mercury »
PDB 1rsv-1yp2 »
1tlf »
Mercury in PDB 1tlf: Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping
Protein crystallography data
The structure of Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping, PDB code: 1tlf
was solved by
A.M.Friedman,
T.O.Fischmann,
T.A.Steitz,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | N/A / 2.60 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 90.160, 64.730, 117.940, 90.00, 91.75, 90.00 |
| R / Rfree (%) | 22.2 / n/a |
Mercury Binding Sites:
The binding sites of Mercury atom in the Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping
(pdb code 1tlf). This binding sites where shown within
5.0 Angstroms radius around Mercury atom.
In total 4 binding sites of Mercury where determined in the Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping, PDB code: 1tlf:
Jump to Mercury binding site number: 1; 2; 3; 4;
In total 4 binding sites of Mercury where determined in the Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping, PDB code: 1tlf:
Jump to Mercury binding site number: 1; 2; 3; 4;
Mercury binding site 1 out of 4 in 1tlf
Go back to
Mercury binding site 1 out
of 4 in the Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping
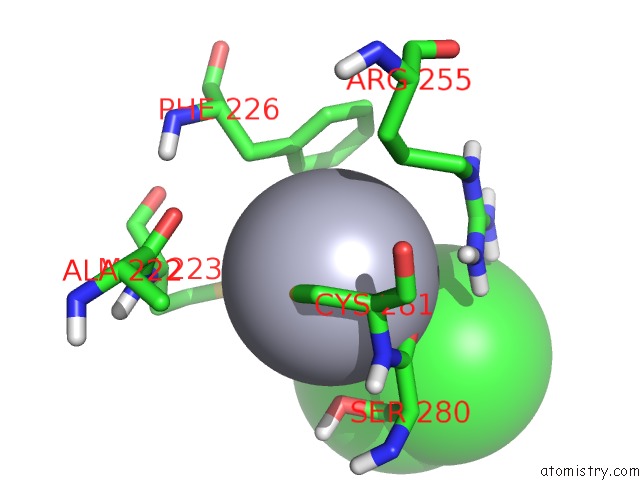
Mono view
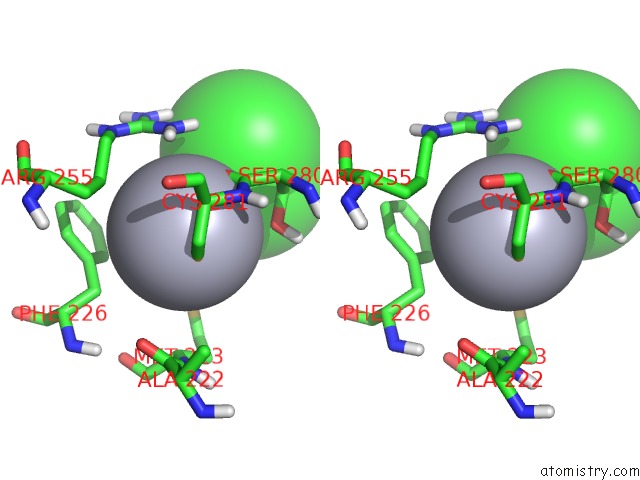
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 1 of Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping within 5.0Å range:
|
Mercury binding site 2 out of 4 in 1tlf
Go back to
Mercury binding site 2 out
of 4 in the Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping
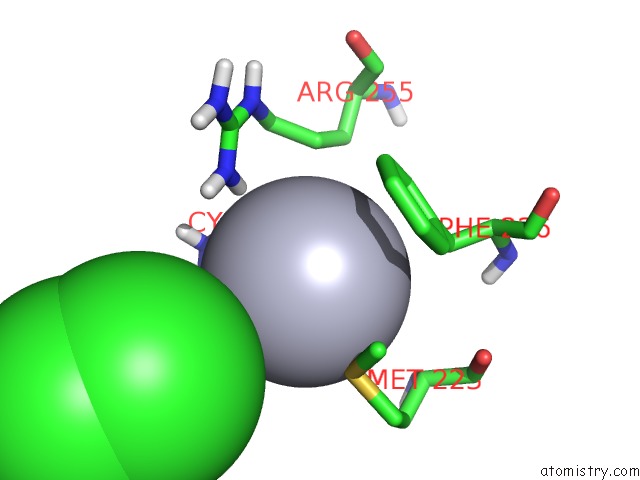
Mono view
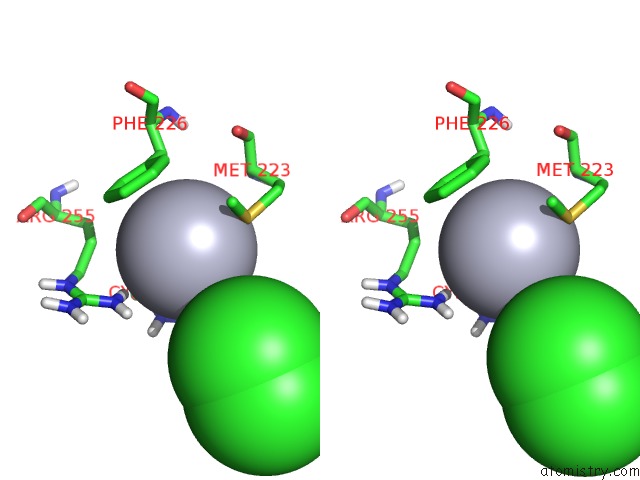
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 2 of Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping within 5.0Å range:
|
Mercury binding site 3 out of 4 in 1tlf
Go back to
Mercury binding site 3 out
of 4 in the Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping
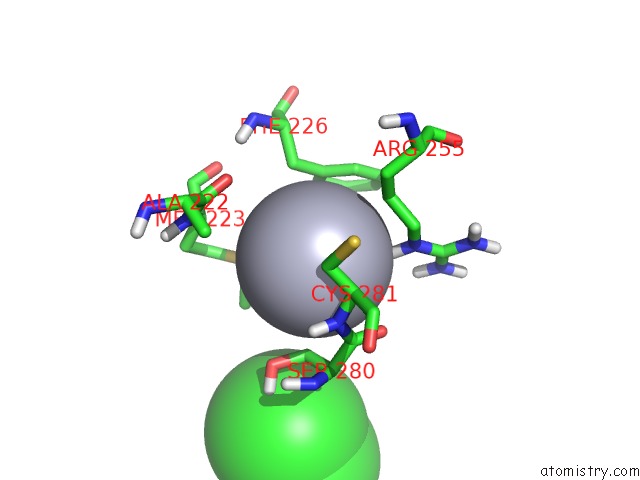
Mono view
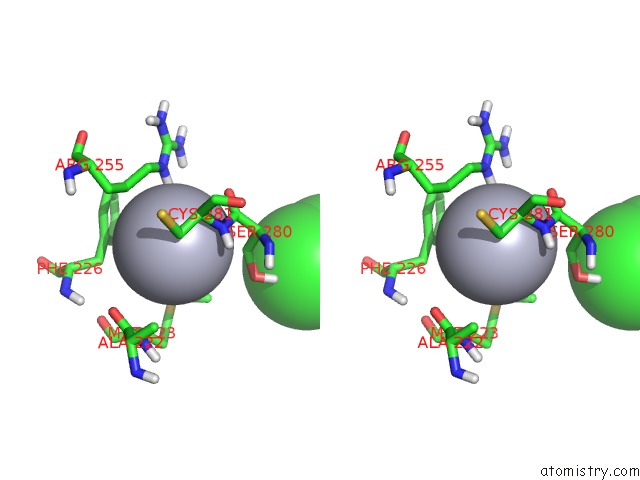
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 3 of Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping within 5.0Å range:
|
Mercury binding site 4 out of 4 in 1tlf
Go back to
Mercury binding site 4 out
of 4 in the Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping
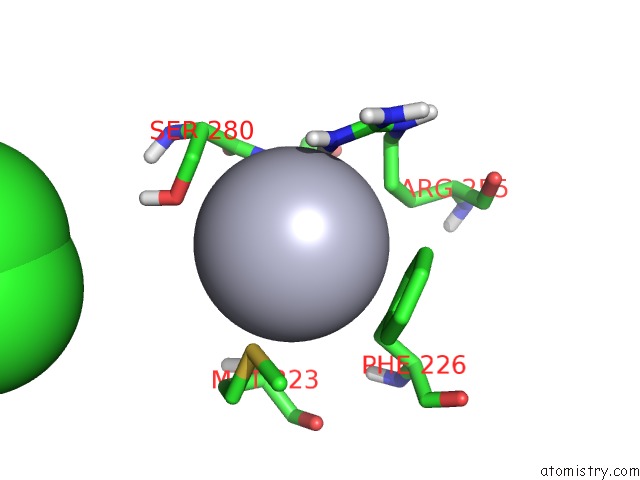
Mono view
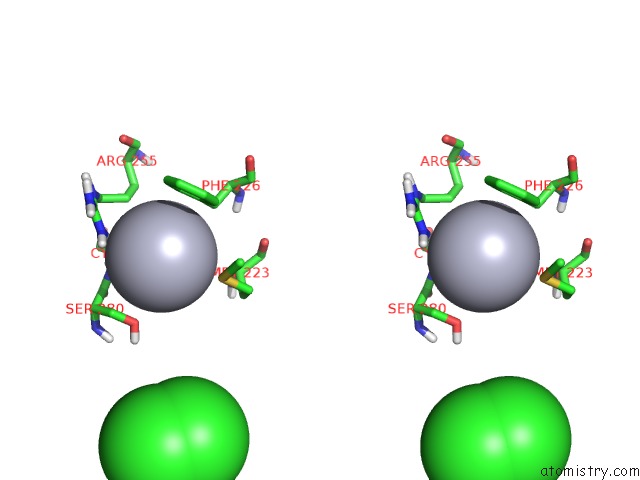
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 4 of Unprecedented Quaternary Structure of E. Coli Lac Repressor Core Tetramer: Implications For Dna Looping within 5.0Å range:
|
Reference:
A.M.Friedman,
T.O.Fischmann,
T.A.Steitz.
Crystal Structure of Lac Repressor Core Tetramer and Its Implications For Dna Looping. Science V. 268 1721 1995.
ISSN: ISSN 0036-8075
PubMed: 7792597
Page generated: Fri Aug 8 09:33:23 2025
ISSN: ISSN 0036-8075
PubMed: 7792597
Last articles
I in 2GJMI in 2GSQ
I in 2GGQ
I in 2FWZ
I in 2G19
I in 2G1M
I in 2DGM
I in 2FWE
I in 2FWF
I in 2FWH