Mercury »
PDB 1rsv-1yp2 »
1ugc »
Mercury in PDB 1ugc: Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H)
Enzymatic activity of Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H)
All present enzymatic activity of Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H):
4.2.1.1;
4.2.1.1;
Protein crystallography data
The structure of Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H), PDB code: 1ugc
was solved by
L.R.Scolnick,
D.W.Christianson,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 6.50 / 2.00 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 42.700, 41.700, 73.000, 90.00, 104.60, 90.00 |
| R / Rfree (%) | 16.2 / 22.5 |
Other elements in 1ugc:
The structure of Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H) also contains other interesting chemical elements:
| Zinc | (Zn) | 1 atom |
Mercury Binding Sites:
The binding sites of Mercury atom in the Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H)
(pdb code 1ugc). This binding sites where shown within
5.0 Angstroms radius around Mercury atom.
In total only one binding site of Mercury was determined in the Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H), PDB code: 1ugc:
In total only one binding site of Mercury was determined in the Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H), PDB code: 1ugc:
Mercury binding site 1 out of 1 in 1ugc
Go back to
Mercury binding site 1 out
of 1 in the Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H)
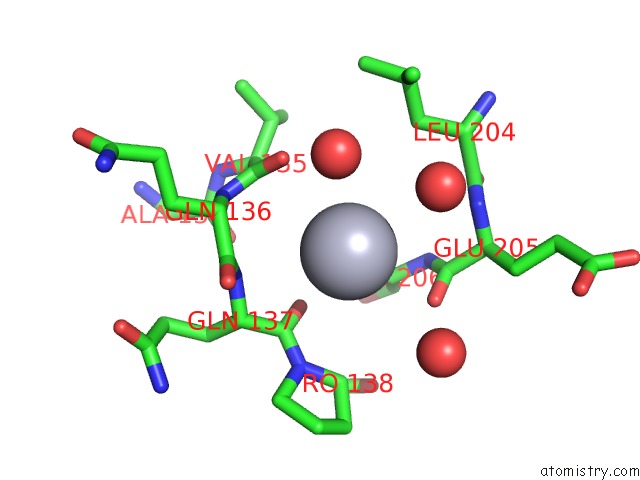
Mono view
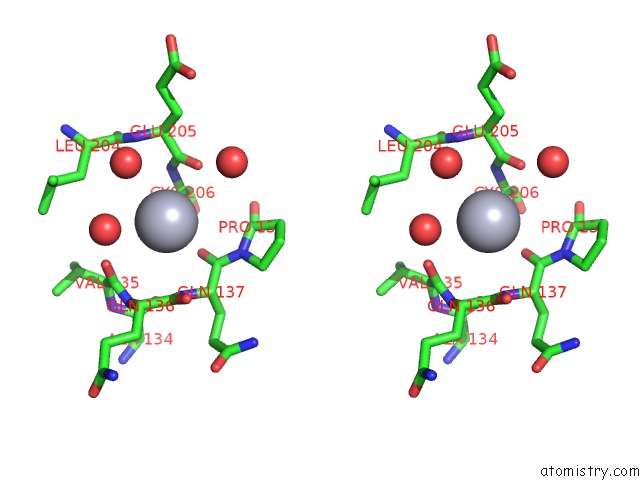
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 1 of Human Carbonic Anhydrase II [Hcaii] (E.C.4.2.1.1) Mutant with Ala 65 Replaced By His (A65H) within 5.0Å range:
|
Reference:
L.R.Scolnick,
D.W.Christianson.
X-Ray Crystallographic Studies of Alanine-65 Variants of Carbonic Anhydrase II Reveal the Structural Basis of Compromised Proton Transfer in Catalysis. Biochemistry V. 35 16429 1996.
ISSN: ISSN 0006-2960
PubMed: 8987974
DOI: 10.1021/BI9617872
Page generated: Fri Aug 8 09:33:34 2025
ISSN: ISSN 0006-2960
PubMed: 8987974
DOI: 10.1021/BI9617872
Last articles
I in 2GJMI in 2GSQ
I in 2GGQ
I in 2FWZ
I in 2G19
I in 2G1M
I in 2DGM
I in 2FWE
I in 2FWF
I in 2FWH