Mercury »
PDB 3kbc-3wa8 »
3px7 »
Mercury in PDB 3px7: Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate
Enzymatic activity of Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate
All present enzymatic activity of Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate:
5.99.1.2;
5.99.1.2;
Protein crystallography data
The structure of Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate, PDB code: 3px7
was solved by
Z.Zhang,
Y.C.Tse-Dinh,
B.Cheng,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 500.00 / 2.30 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 61.602, 91.759, 141.976, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 23 / 26.9 |
Mercury Binding Sites:
The binding sites of Mercury atom in the Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate
(pdb code 3px7). This binding sites where shown within
5.0 Angstroms radius around Mercury atom.
In total 3 binding sites of Mercury where determined in the Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate, PDB code: 3px7:
Jump to Mercury binding site number: 1; 2; 3;
In total 3 binding sites of Mercury where determined in the Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate, PDB code: 3px7:
Jump to Mercury binding site number: 1; 2; 3;
Mercury binding site 1 out of 3 in 3px7
Go back to
Mercury binding site 1 out
of 3 in the Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate
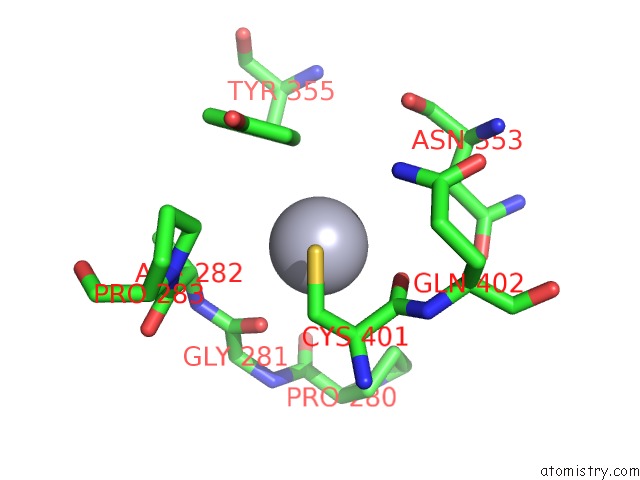
Mono view
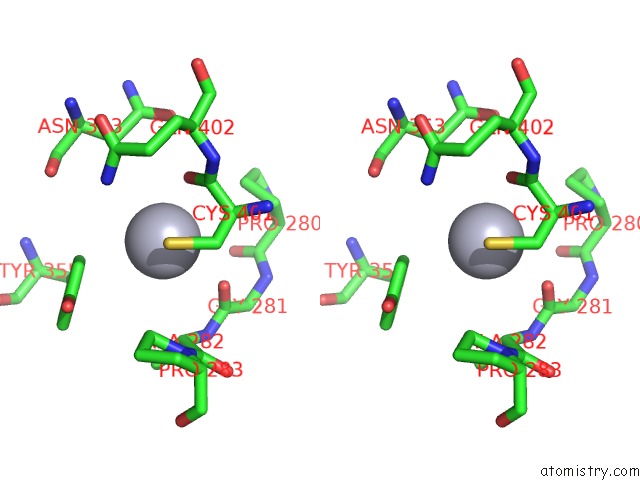
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 1 of Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate within 5.0Å range:
|
Mercury binding site 2 out of 3 in 3px7
Go back to
Mercury binding site 2 out
of 3 in the Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate
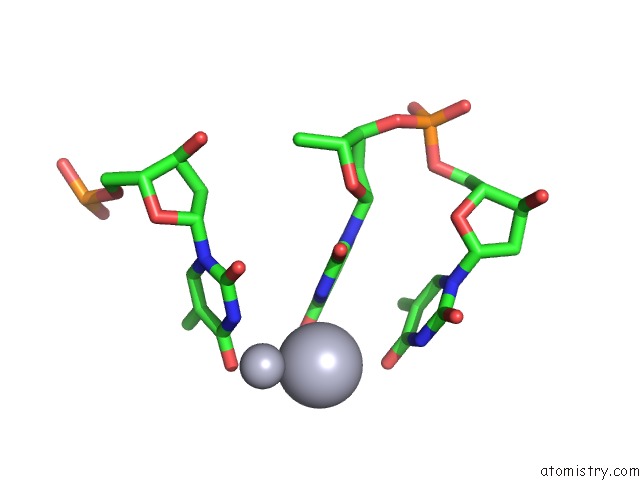
Mono view
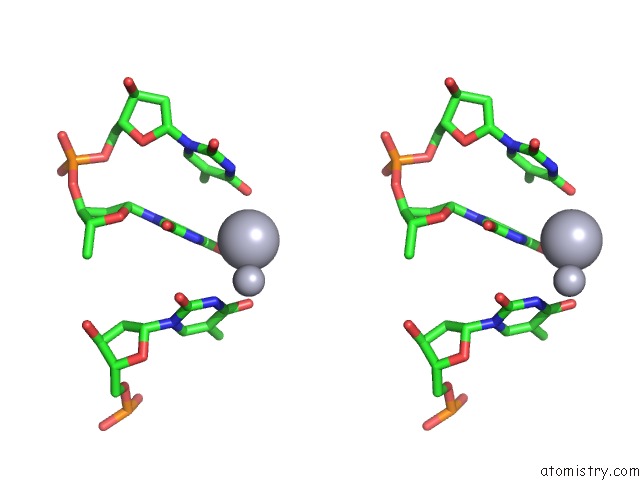
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 2 of Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate within 5.0Å range:
|
Mercury binding site 3 out of 3 in 3px7
Go back to
Mercury binding site 3 out
of 3 in the Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate
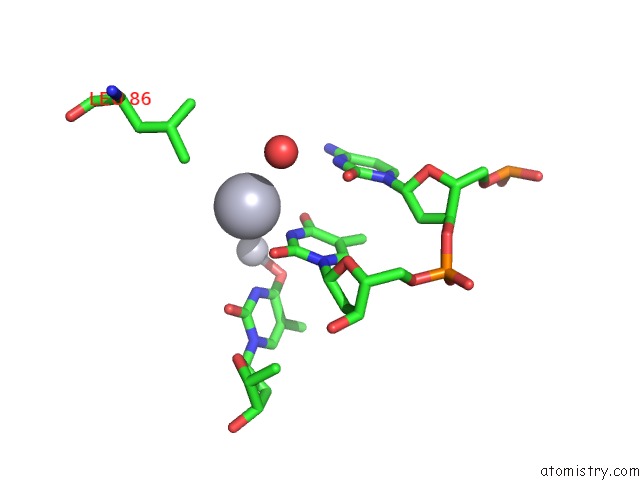
Mono view
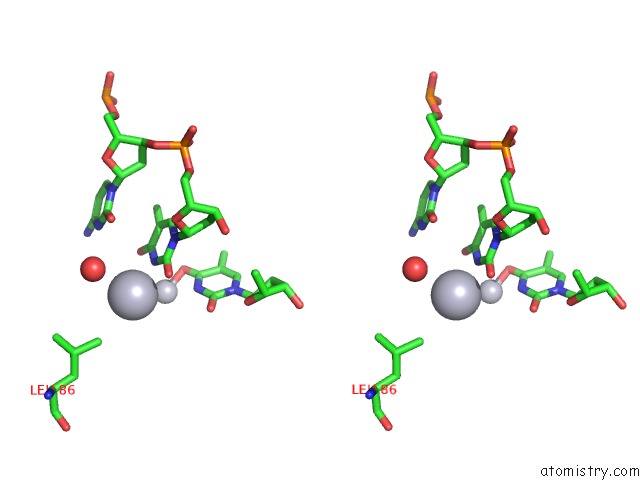
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 3 of Crystal Structure of Covalent Complex of Topoisomerase 1A with Substrate within 5.0Å range:
|
Reference:
Y.C.Tse-Dinh,
Z.Zhang,
B.Cheng.
Structure of E. Coli Topoisomerase I Trapped As Covalent Complex Intermediate with Cleaved Dna To Be Published.
Page generated: Sun Aug 11 04:07:44 2024
Last articles
Cl in 5RAVCl in 5RAU
Cl in 5RAS
Cl in 5RAR
Cl in 5RAQ
Cl in 5RAP
Cl in 5RAO
Cl in 5RAN
Cl in 5RAJ
Cl in 5RAM