Mercury »
PDB 3wee-4ia4 »
4fyf »
Mercury in PDB 4fyf: Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase
Enzymatic activity of Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase
All present enzymatic activity of Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase:
3.1.3.67;
3.1.3.67;
Protein crystallography data
The structure of Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase, PDB code: 4fyf
was solved by
F.S.Hsu,
W.Zhu,
L.Brennan,
L.Tao,
Z.Q.Luo,
Y.Mao,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 47.18 / 2.42 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 71.835, 115.709, 125.114, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 21.1 / 27 |
Mercury Binding Sites:
The binding sites of Mercury atom in the Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase
(pdb code 4fyf). This binding sites where shown within
5.0 Angstroms radius around Mercury atom.
In total 3 binding sites of Mercury where determined in the Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase, PDB code: 4fyf:
Jump to Mercury binding site number: 1; 2; 3;
In total 3 binding sites of Mercury where determined in the Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase, PDB code: 4fyf:
Jump to Mercury binding site number: 1; 2; 3;
Mercury binding site 1 out of 3 in 4fyf
Go back to
Mercury binding site 1 out
of 3 in the Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase
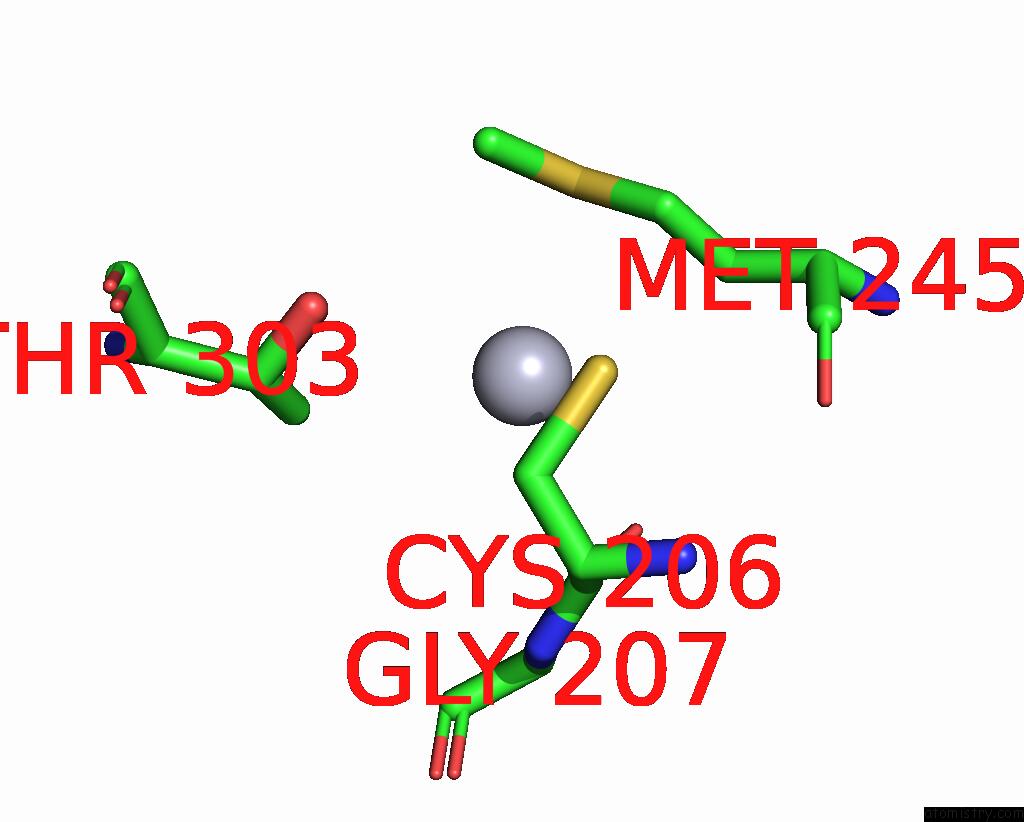
Mono view
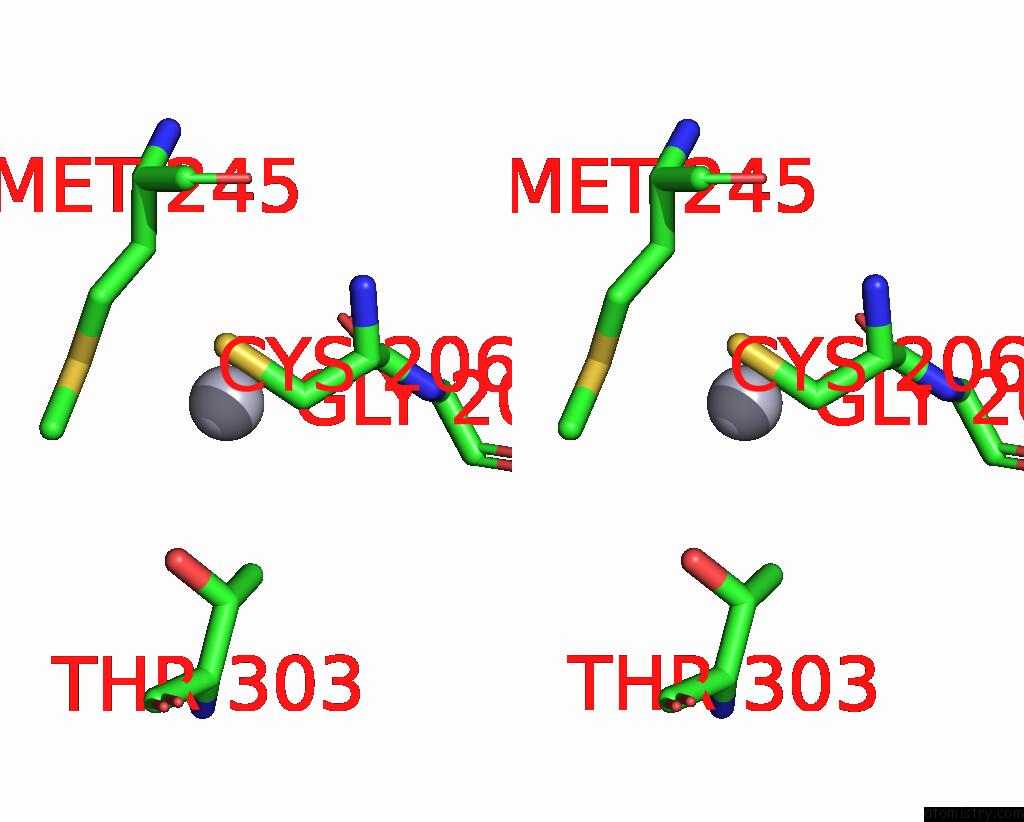
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 1 of Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase within 5.0Å range:
|
Mercury binding site 2 out of 3 in 4fyf
Go back to
Mercury binding site 2 out
of 3 in the Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase
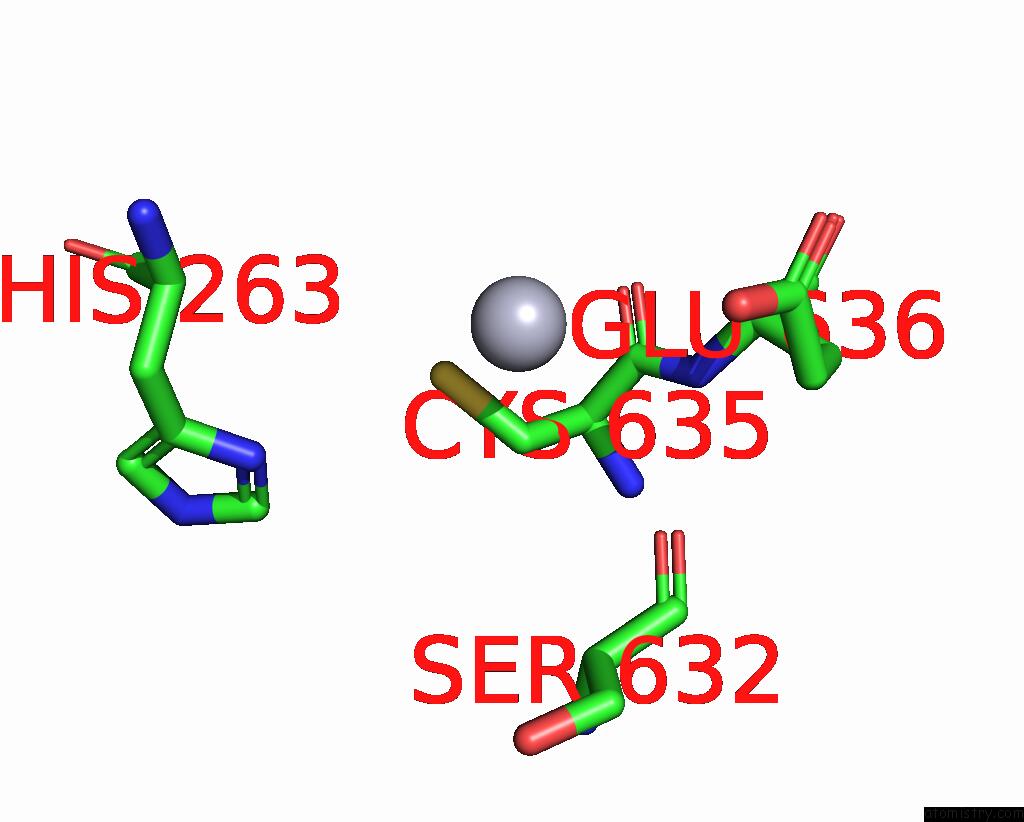
Mono view
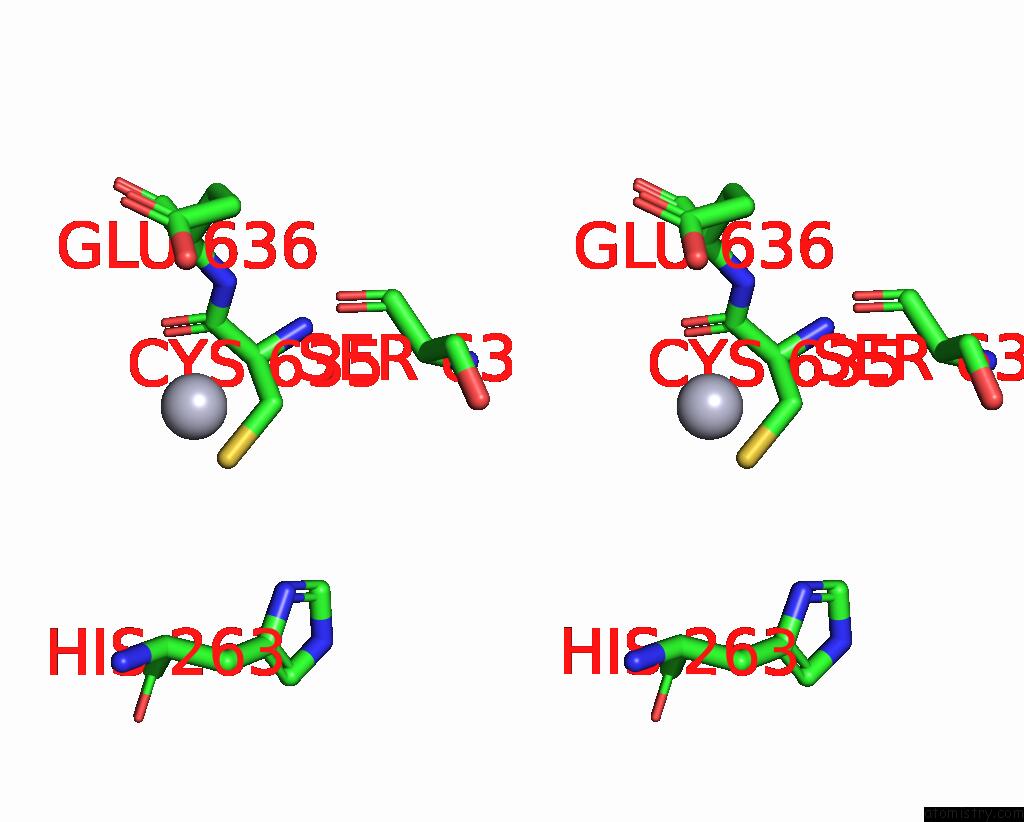
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 2 of Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase within 5.0Å range:
|
Mercury binding site 3 out of 3 in 4fyf
Go back to
Mercury binding site 3 out
of 3 in the Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase

Mono view
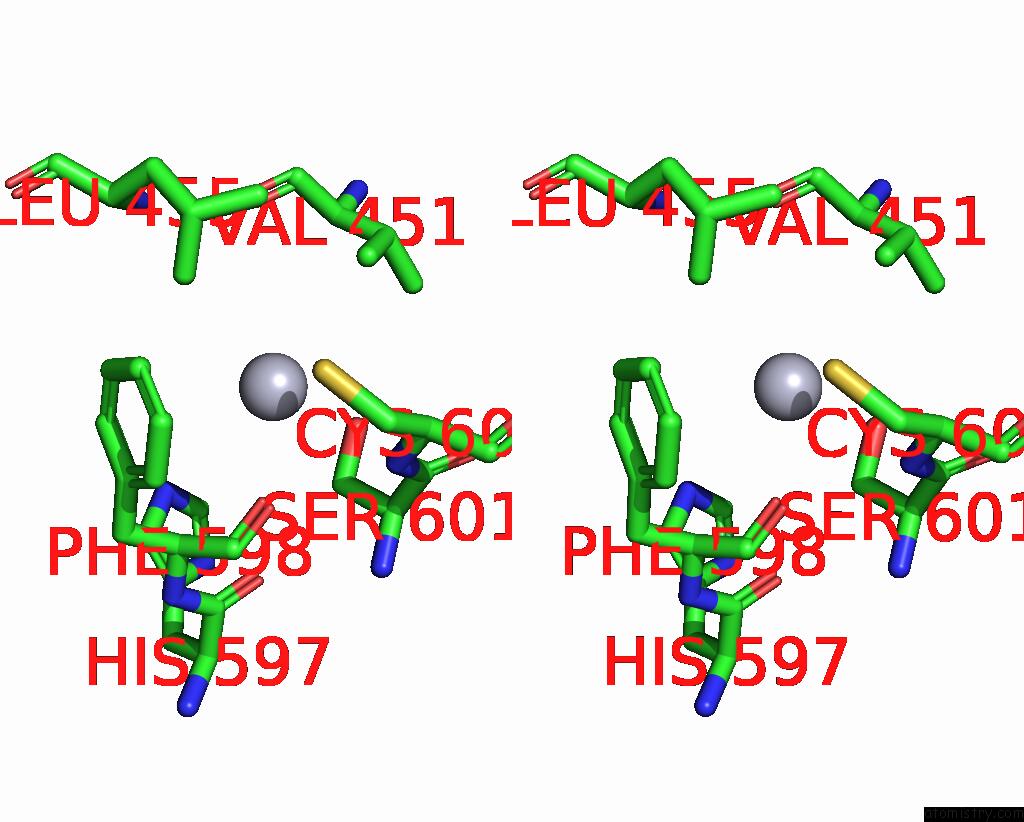
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 3 of Structural Basis For Substrate Recognition By A Novel Legionella Phosphoinositide Phosphatase within 5.0Å range:
|
Reference:
F.Hsu,
W.Zhu,
L.Brennan,
L.Tao,
Z.Q.Luo,
Y.Mao.
Structural Basis For Substrate Recognition By A Unique Legionella Phosphoinositide Phosphatase. Proc.Natl.Acad.Sci.Usa V. 109 13567 2012.
ISSN: ISSN 0027-8424
PubMed: 22872863
DOI: 10.1073/PNAS.1207903109
Page generated: Sun Aug 11 04:41:33 2024
ISSN: ISSN 0027-8424
PubMed: 22872863
DOI: 10.1073/PNAS.1207903109
Last articles
Cl in 7TBVCl in 7T8Q
Cl in 7TCD
Cl in 7TB0
Cl in 7TBS
Cl in 7TBQ
Cl in 7TAZ
Cl in 7TAY
Cl in 7TAV
Cl in 7T8P