Mercury »
PDB 4q81-5c9l »
5c17 »
Mercury in PDB 5c17: Crystal Structure of the Mercury-Bound Form of MERB2
Protein crystallography data
The structure of Crystal Structure of the Mercury-Bound Form of MERB2, PDB code: 5c17
was solved by
H.M.Wahba,
L.Lecoq,
M.Stevenson,
A.Mansour,
L.Cappadocia,
J.Lafrance-Vanasse,
K.J.Wilkinson,
J.Sygusch,
D.E.Wilcox,
J.G.Omichinski,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.46 / 1.24 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 83.290, 53.421, 47.719, 90.00, 96.85, 90.00 |
| R / Rfree (%) | 14.2 / 16.1 |
Mercury Binding Sites:
The binding sites of Mercury atom in the Crystal Structure of the Mercury-Bound Form of MERB2
(pdb code 5c17). This binding sites where shown within
5.0 Angstroms radius around Mercury atom.
In total only one binding site of Mercury was determined in the Crystal Structure of the Mercury-Bound Form of MERB2, PDB code: 5c17:
In total only one binding site of Mercury was determined in the Crystal Structure of the Mercury-Bound Form of MERB2, PDB code: 5c17:
Mercury binding site 1 out of 1 in 5c17
Go back to
Mercury binding site 1 out
of 1 in the Crystal Structure of the Mercury-Bound Form of MERB2
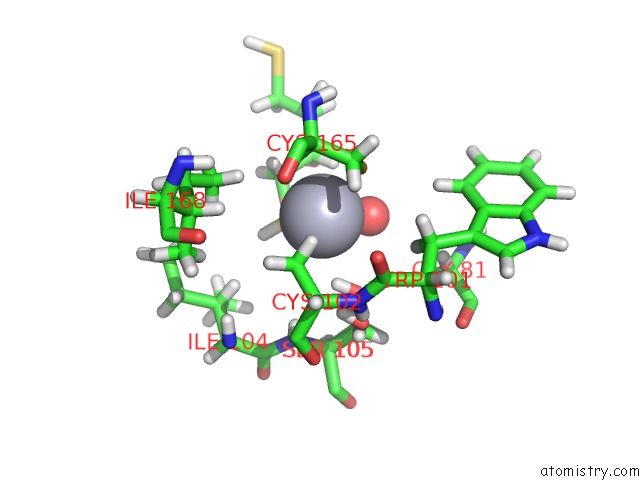
Mono view
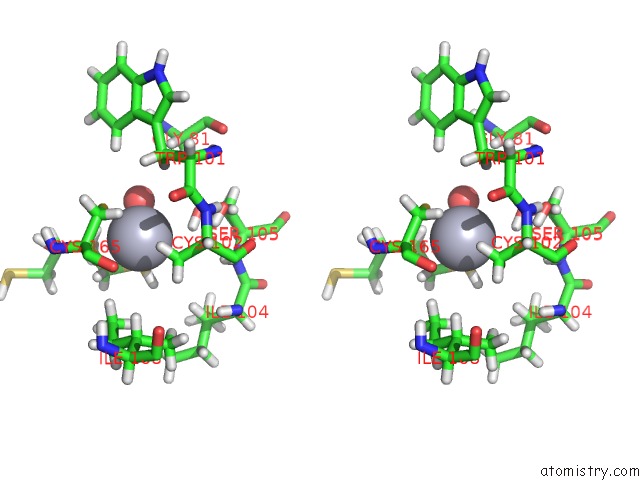
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 1 of Crystal Structure of the Mercury-Bound Form of MERB2 within 5.0Å range:
|
Reference:
H.M.Wahba,
L.Lecoq,
M.Stevenson,
A.Mansour,
L.Cappadocia,
J.Lafrance-Vanasse,
K.J.Wilkinson,
J.Sygusch,
D.E.Wilcox,
J.G.Omichinski.
Structural and Biochemical Characterization of A Copper-Binding Mutant of the Organomercurial Lyase Merb: Insight Into the Key Role of the Active Site Aspartic Acid in Hg-Carbon Bond Cleavage and Metal Binding Specificity. Biochemistry V. 55 1070 2016.
ISSN: ISSN 0006-2960
PubMed: 26820485
DOI: 10.1021/ACS.BIOCHEM.5B01298
Page generated: Sun Aug 11 05:41:22 2024
ISSN: ISSN 0006-2960
PubMed: 26820485
DOI: 10.1021/ACS.BIOCHEM.5B01298
Last articles
Ca in 5S5UCa in 5S5T
Ca in 5S5S
Ca in 5S5R
Ca in 5S5Q
Ca in 5S5P
Ca in 5S5O
Ca in 5S5N
Ca in 5S5M
Ca in 5S5L