Mercury »
PDB 7tfx-8p6u »
8dx1 »
Mercury in PDB 8dx1: [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops
Protein crystallography data
The structure of [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops, PDB code: 8dx1
was solved by
B.Lu,
S.Vecchioni,
N.C.Seeman,
R.Sha,
Y.P.Ohayon,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 39.72 / 4.79 |
| Space group | H 3 |
| Cell size a, b, c (Å), α, β, γ (°) | 106.18, 106.18, 88.077, 90, 90, 120 |
| R / Rfree (%) | 24.5 / 28.9 |
Other elements in 8dx1:
The structure of [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops also contains other interesting chemical elements:
| Silver | (Ag) | 1 atom |
Mercury Binding Sites:
The binding sites of Mercury atom in the [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops
(pdb code 8dx1). This binding sites where shown within
5.0 Angstroms radius around Mercury atom.
In total only one binding site of Mercury was determined in the [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops, PDB code: 8dx1:
In total only one binding site of Mercury was determined in the [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops, PDB code: 8dx1:
Mercury binding site 1 out of 1 in 8dx1
Go back to
Mercury binding site 1 out
of 1 in the [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops
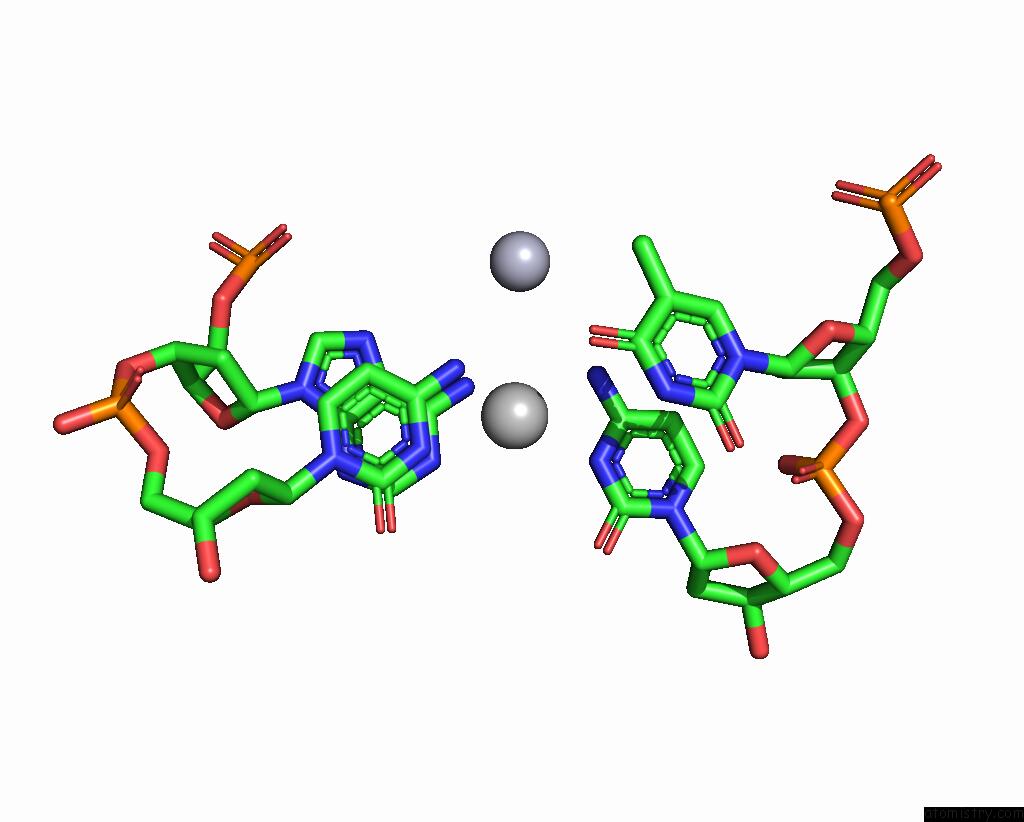
Mono view
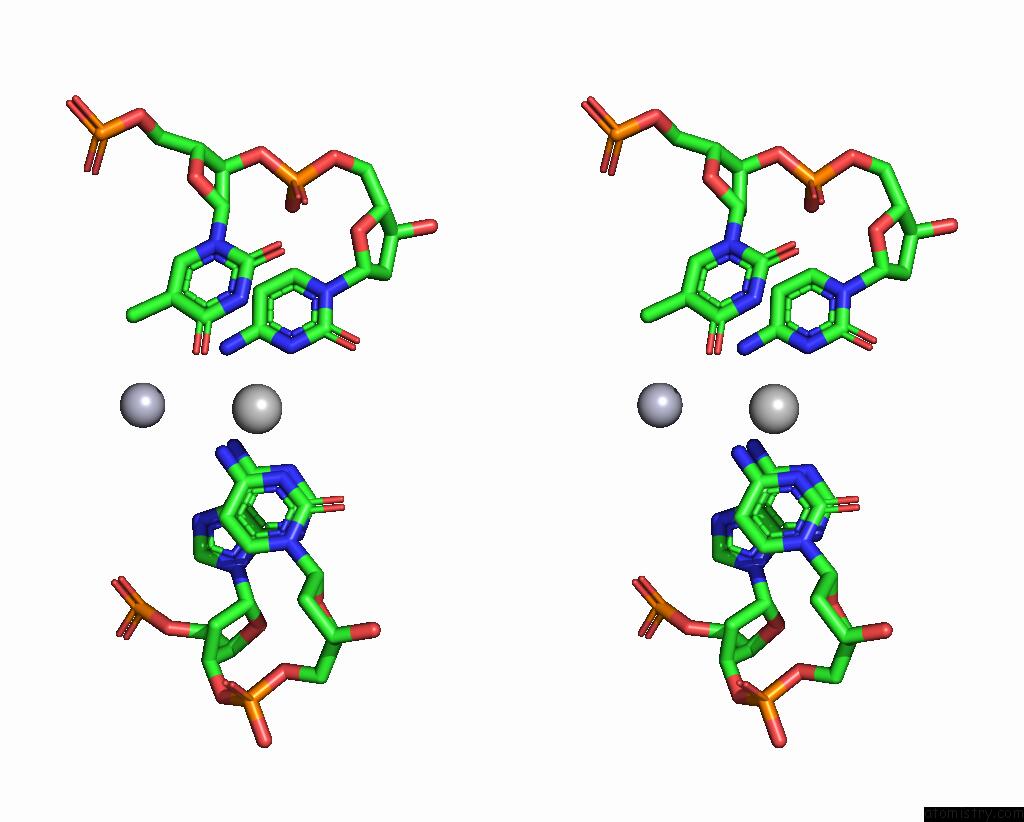
Stereo pair view

Mono view

Stereo pair view
A full contact list of Mercury with other atoms in the Hg binding
site number 1 of [C:HG2+/Ag+:C--pH 7 Mops] Metal-Mediated Dna Base Pair in Tensegrity Triangle in Ag+ and HG2+ Solution in Mops within 5.0Å range:
|
Reference:
B.Lu,
Y.P.Ohayon,
K.Woloszyn,
C.F.Yang,
J.B.Yoder,
L.J.Rothschild,
S.J.Wind,
W.A.Hendrickson,
C.Mao,
N.C.Seeman,
J.W.Canary,
R.Sha,
S.Vecchioni.
Heterobimetallic Base Pair Programming in Designer 3D Dna Crystals. J.Am.Chem.Soc. 2023.
ISSN: ESSN 1520-5126
PubMed: 37530628
DOI: 10.1021/JACS.3C05478
Page generated: Sun Aug 11 08:56:49 2024
ISSN: ESSN 1520-5126
PubMed: 37530628
DOI: 10.1021/JACS.3C05478
Last articles
Cl in 2VYXCl in 2W0R
Cl in 2W0D
Cl in 2VX3
Cl in 2VZ5
Cl in 2VYO
Cl in 2VYA
Cl in 2VY3
Cl in 2VY0
Cl in 2VXT